서론
재료 및 방법
식물 및 바이러스
바이러스 증식 및 주형 준비
검정용 프라이머 분석 및 제작
Table 1
RT-PCR 반응액과 증폭산물 확인
RT-PCR mix와 프라이머 조합별 특이도 비교
RT-PCR mix 종류와 주형농도, cDNA 합성온도 및 시간별 반응양상 비교
선발 프라이머 검출감도 및 특이도 비교
결과
프라이머 조합별 특이도 비교
Table 2
| Virus | Code | Forward primer | Reverse primer | Amplicon size (bp) | Nonspecific background product* |
|---|---|---|---|---|---|
| ToCV | C029 | ATCGGACATTATGTTCAAGG | AAATCCTCATTACCACATCG | 281 | Weak |
| C030 | CCTGATTGGTTCTAAACTGC | CCACACCTACCACGATACTT | 198 | Strong | |
| C031 | ATTGCTTAGACCGATTCAGA | TCACCCATTTTAGCGTACTT | 372 | Strong | |
| C032 | TATGTGTCAGGCCATTGTAA | TTCATAAGCAGGTTCGAGAT | 348 | Weak | |
| C033 | TGATGTTTCCAGTGTTTTCA | TGGTAAATGAAATTCCCAAC | 418 | Strong | |
| C034 | GTGTTACGATGGGTTTGTTT | CTTGAGTTTGTCCCAAAGAG | 559 | Strong | |
| C035 | TTCCAAGTTCGATAGGAAGA | CGAAACATATAACGACGAATC | 565 | Strong | |
| C036 | TCGACTTTTCGAGGTTGTAT | CCACACCTACCACGATACTT | 556 | Strong | |
| C037 | CATGTTTTCCAACAGGAGAT | GGGATTATCATATTGCCTCA | 832 | Strong | |
| C038 | CTCTTTGGGACAAACTCAAG | ATCGAACTTGGAAACGAGTA | 980 | Strong | |
| C039 | GATGAAACCGAAGATGGATA | AACAACAGTCAAATCGCTCT | 943 | Strong | |
| C040 | TCTGATGCCTTGGTTACTTT | AAGCAACAAACCTACCTTGA | 564 | Weak | |
| C041 | TGATGTTTCCAGTGTTTTCA | TAGTTTGCACCAAAGGAACT | 527 | No | |
| C042 | ATCTCGAACCTGCTTATGAA | TCAGACTCGGGTTAGGTAAA | 657 | Strong | |
| C043 | TTTACCTAACCCGAGTCTGA | TAGTTTGCACCAAAGGAACT | 713 | Strong | |
| C044 | TACAGCTTAACCCGTTGAAT | CATTGTCAATACAGTGGCAG | 518 | No | |
| C045 | TAGACACATGGCAATGGTAA | AAAACGCACTGTCTGTACCT | 1,055 | Strong | |
| PepMoV | C046 | GATGGATTTGGCTACAACAT | CAGCTAATTCAAAAGCCCTA | 443 | Strong |
| C047 | AATAGTGTGTTTGGGTCGTC | CTACCTGTGCCTTGACTCTC | 308 | Strong | |
| C048 | TGTTCACTAGGCTCAGGAGT | GACGACCCAAACACACTATT | 461 | Weak | |
| C049 | GTGATGAATGGCTTAATGGT | CCAAATAACCTTGTTTGAGC | 371 | Strong | |
| C050 | ACACTGGCAATAATGCTTTT | TCCAGTGGATTTACCAGAAC | 390 | Strong | |
| C051 | TATCTGGTGGTCCATTTTTC | CTACCTGTGCCTTGACTCTC | 355 | Strong | |
| C052 | TGTTCACTAGGCTCAGGAGT | GAAAAATGGACCACCAGATA | 414 | Strong | |
| C053 | GTGGTCTCTGGGATGAAATA | CCAACTACGATCACCTCCT | 249 | Strong | |
| C054 | GAATGCTAGTTTTCTGTGGG | ATGTTGTAGCCAAATCCATC | 557 | Weak | |
| C055 | CACCATTCCAAGAATCAAAT | CCAAATAACCTTGTTTGAGC | 576 | Weak | |
| C056 | ATGATGAAAAACAGGTCGTC | GACGACCCAAACACACTATT | 613 | Strong | |
| C057 | AAGAGGAGCGAATTGTATCA | ACCATTAAGCCATTCATCAC | 694 | Strong | |
| C058 | GTTTTGCTGATAGAACCGAC | TGAATTCGTTCTCCGTAACT | 740 | Strong | |
| C059 | GAAGCTGCTCTGTATTGCTT | CAAGAGAAGACTTGGACTGG | 769 | Strong | |
| C060 | GACGCTCTAAGACGAAAAGA | ATCAGGTTTGGCACTCTAAA | 772 | Strong | |
| C061 | AAAGAATTCAGGCATTGAAA | ATCAGGTTTGGCACTCTAAA | 757 | Strong | |
| C062 | GGCTCTAAAAGCTGAACTGA | TGATACAATTCGCTCCTCTT | 745 | Strong | |
| C063 | AGTGGTTGGAATCTCAAATG | CAGGGAATTTCATTATAGCG | 815 | Strong | |
| ToMV | C064 | TAGTTCAGCAAAGGTGGTTT | ACGTCTGCATAAGTCTCACC | 681 | Strong |
| C065 | TTCACACAGTCTGACAAGGA | GGAAGATCCACAAAATCAAA | 824 | Weak | |
| C066 | AGATTGCTCAAAGATGGAGA | AATTTCATGGTAACAGCGTC | 1,064 | Weak | |
| CMV | C067 | CAGACTTATTCGTACCGAGG | GTAGATGATTTCGCGGATAG | 323 | Strong |
| C068 | ACTCTCTGACGAGTTCGGTA | CAATTCCATAATTCTTTCGC | 392 | Strong | |
| C069 | GTCGAGTCATGGACAAATCT | ACTGACCATTTTAGCCGTAA | 803 | Weak | |
| C070 | ACATCAATGAATTGGTAGCC | CTCGGTACGAATAAGTCTGG | 876 | Weak | |
| C071 | CTATCCGCGAAATCATCTAC | TCAACTTGACTAATGGTCCC | 959 | Weak | |
| TSWV | C072 | GCAATTCTGGTTCTCTATGC | CATTCTTCAGAGTGTCCCAT | 375 | Weak |
| C073 | CTAATGAAATCGGAAAATCG | AGAATAATCGGATAGTCGCA | 197 | No | |
| C074 | GACAGCCTGGAACATAAAAG | GAAAATGTGGAACACAAGGT | 673 | Strong |
ToCV, Tomato chlorosis virus; PepMoV, Pepper mottle virus; ToMV, Tomato mosaic virus; CMV, Cucumber mosaic virus; TSWV, Tomato spotted wilt virus.
* RT-PCR was performed with each primer combination in the RT-PCR mix3 without template RNA. The condition was as follows cDNA synthesis (42°C/20 min), denaturation (95°C/10 min), amplification (35 cycles of 95°C/1 min, 58.5°C/1 min, 60.7°C/1 min, 62.4°C/1 min, and 72°C/1 min), and final extension (72°C/10 min). The result was determined by visual observation for the average amount or intensity of nonspecific background products of each gel.
진단용 RT-PCR mix 선발
Fig. 1
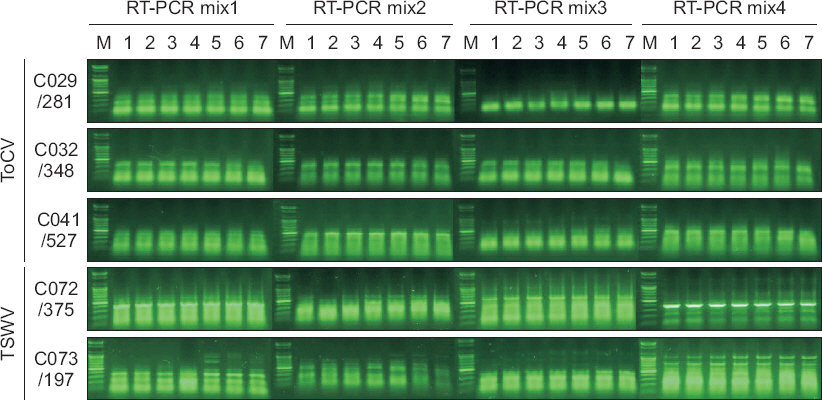
Fig. 2
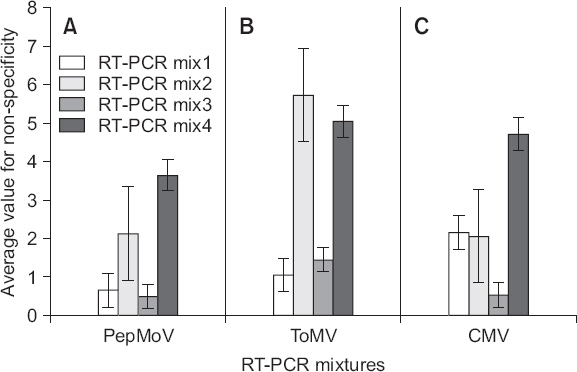
cDNA 합성 조건 최적화 및 선발 프라이머 적용
Fig. 3
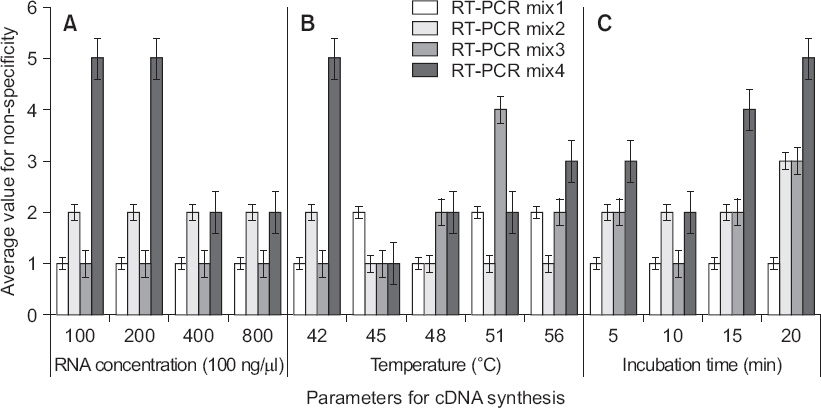
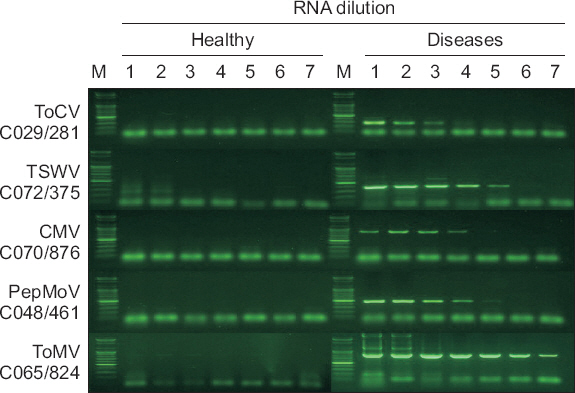
Fig. 4



 PDF Links
PDF Links PubReader
PubReader Full text via DOI
Full text via DOI Download Citation
Download Citation Print
Print






