Sclerotinia sclerotiorum is a devastating, necrotrophic, and cosmopolitan plant pathogen with a broad host range (
Boland and Hall, 1994;
Bolton et al., 2005). In Korea, 60 plant species have been reported as hosts of
S. sclerotiorum (
Kim et al., 2009). The optimal temperature range for
Sclerotinia growth is from 15-21┬░C, and cool and moist conditions increase the incidence of field rot. Green peas have a short season and a limited shelf-life, so preservation ensures long-term availability. Growth can also occur in lower temperatures (0-4┬░C); therefore, green peas are also susceptible to fungal growth during transit and storage (
Naito and Sugimoto, 1986;
Willetts and Wong, 1980). This study identifies
S. sclerotiorum as the causal agent of a post-harvest green pea pod rot based on morphological characteristics, phylogenetic analysis, and a pathogenicity test.
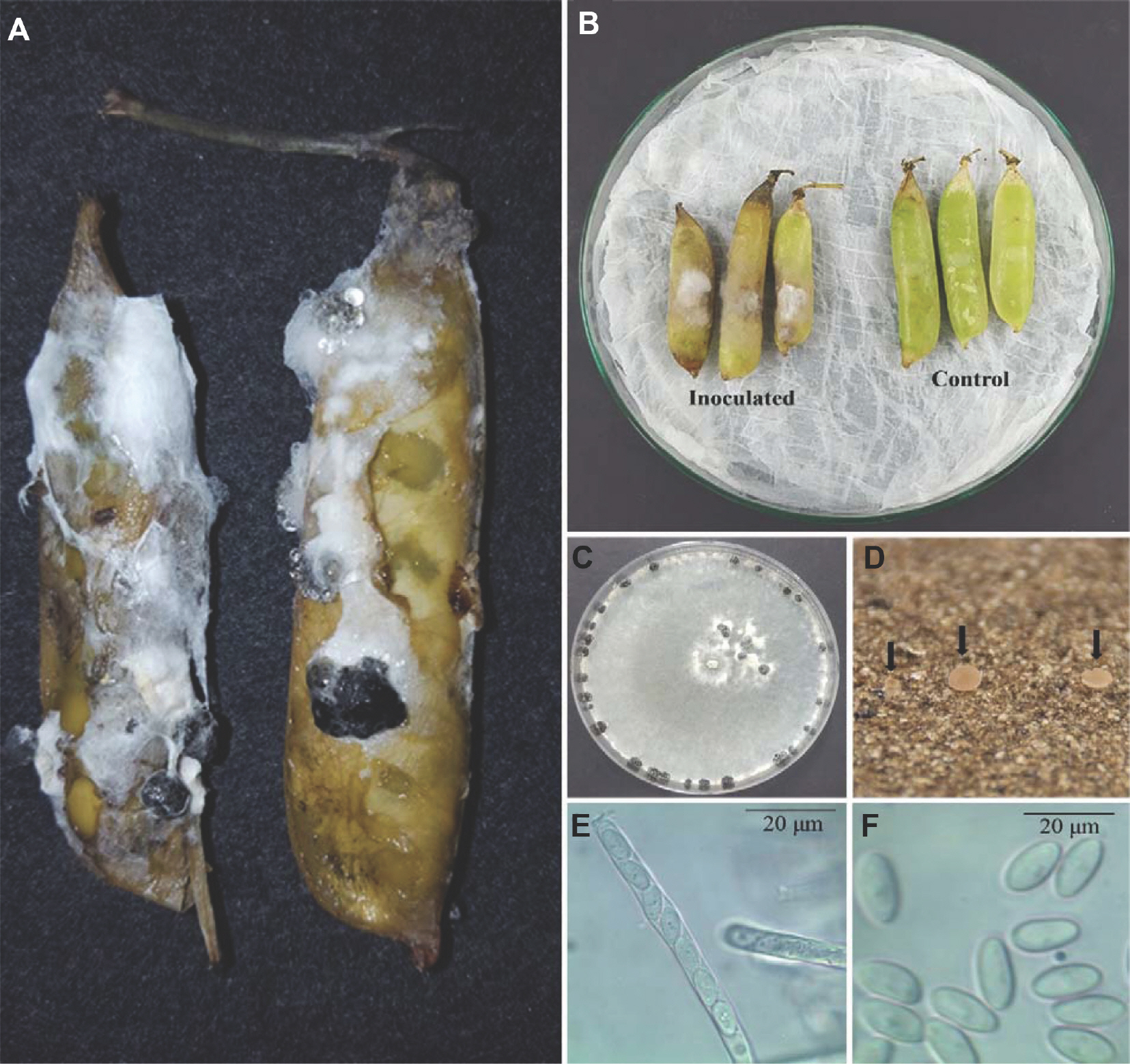
Green pea pods harvested from Gangneung, Gangwon Province, South Korea (June 2017) exhibited post-harvest rot symptoms (white and fuzzy mycelial growth with black sclerotia) after 5-7 days of refrigeration (4┬░C, 70% relative humidity) (
Fig. 1A). The pathogen was isolated by excising the infected pod tissue and then surface-sterilized in 1% sodium hypochlorite (NaOCl) for one minute. The tissue was rinsed with sterilized distilled water (3├Ś), and then plated on potato dextrose agar (PDA; Difco, Detroit, MI, USA). After 4 days of room temperature (RT; 20┬▒2┬░C) incubation, the culture was purified by the hyphal-tip method and main-tained on PDA plates. Five morphologically similar fungal isolates were obtained from five green pea pod rot samples, including isolate GPSS003, which was further examined. The fungus was incubated on PDA medium at RT for 7 days and produced white-gray colonies, and after 3 weeks, showed small sclerotia on the plate peripheries (
Fig. 1C). The sclerotia were black, globosely, cylindrically, or irregularly shaped and between 1.0-7.3├Ś1.2-5.2 mm in size (
n=20).
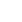
Fig.┬Ā1.
(A) Rotting post-harvest green peas indicated by the fuzzy growths of mycelium and black sclerotia on the pod. (B) Green peas inoculated with Sclerotinia sclerotiorum developed pod rot symptoms after 5 days. (C) S. sclerotiorum three-week-old colonies and black sclerotia growing on potato dextrose agar medium. (D) Apothecia (arrows). (E) Ascus containing eight ascospores. (F) Ascospores.
Sclerotial germination was investigated by placing 10-15, 21-day-old sclerotia in sand Petri dishes. The sclerotia were washed, dried, and pressed into the sand (Ōł╝2 mm depth). The dishes were incubated at RT with alternating 12-hr fluorescent light cycles and were watered three times per week with sterilized water until apothecia developed (
Huang and Kozub, 1989;
Smith and Boland, 1989). A dissecting microscope was used to cut the mature apothecia into thin slices, and the asci and ascospores were examined and photo-graphed with a compound microscope (40├Ś magnification). Sclerotia sand germination generated apothecia after 5-6 weeks of incubation at RT (
Fig. 1D). The apothecia were cup-shaped with a yellowish-brown color and measured 0.6-1.8 cm (
n=4). Asci from the apothecia were cylindrical and had eight spores ranging from 101.2-152.8├Ś5.1-8.5 ╬╝m in size (
n=10) (
Fig. 1E). The ascospores were uniseriate, single-celled, hyaline, and ellipsoid, and between 8.2-13.6├Ś4.0-5.8 ╬╝m in size (
n=30) (
Fig. 1F). These characteristics are con-sistent with the description of
Sclerotinia sclerotiorum (Lib.) de Bary (
Chang and Kim, 2003;
Kohn, 1979) and summarized in
Table 1. A representative isolate (GPSS003) was deposited in the Korean Agricultural Culture Collection, National Institute of Agricultural Science, Rural Development Administration, Wanju, South Korea (KACC48672), for future studies.
Table┬Ā1.
The mycological characteristics of the green pea isolated pathogen and Sclerotinia sclerotiorum
|
Characteristic
|
Present isolate
|
S. sclerotiorum a
|
|
Colony |
|
ŌĆāColor |
White-gray |
White-gray |
|
Sclerotium |
|
ŌĆāShape |
Initially cushion-like, globular, or irregular; dark brown then black |
Initially cushion-like or short-cylindrical; white then black |
|
ŌĆāLength (mm) |
1.0-7.3 |
- |
|
ŌĆāWidth (mm) |
1.2-5.2 |
- |
|
Apothecium |
|
ŌĆāShape |
Cup-shaped with yellowish-brown coloring |
Cup-shaped with yellowish-brown coloring |
|
ŌĆāSize (cm) |
0.6-1.8 |
0.5-2.0 |
|
Ascus |
|
ŌĆāShape |
Cylindrical, hyaline, 8-spored |
Cylindrical, hyaline, 8-spored |
|
ŌĆāLength (╬╝m) |
101.2-152.8 |
110-160.0 |
|
ŌĆāWidth (╬╝m) |
5.1-8.5 |
6.0-10.0 |
|
Ascospore |
|
ŌĆāShape |
Hyaline, ellipsoid to ovoid |
Hyaline, ellipsoid to ovoid |
|
ŌĆāLength (╬╝m) |
8.2-13.6 |
9.0-14.0 |
|
ŌĆāWidth (╬╝m) |
4.0-5.8 |
4.0-6.0 |
For molecular identification, the GPSS003
S. sclerotiorum isolate was grown in potato dextrose broth for 5 days at RT in a shaking incubator (150 rpm). The mycelia were filtered through Whatman No. 1 filter paper, and genomic DNA was extracted using the DNeasy Plant Mini Kit (Qiagen Inc., Valencia, CA, USA). The DNA sequences of the ribosomal DNA internal transcribed spacers (ITS) region (ITS-5.8S rDNA) and beta-tubulin gene (
╬▓tub) were amplified by polymerase chain reaction (PCR) on a Mastercycler Gradient (Eppendorf, Hamburg, Germany) using primers for the ITS region (ITS1, 5ŌĆÖ-TCCGTAGGTGAACCTGCGG-3ŌĆÖ; ITS4, 5ŌĆÖ-TCCTCCGCTTATT-GATATGC-3ŌĆÖ); and the
╬▓tub gene (Bt1a, 5ŌĆÖ-TTCCCCCGTCTC-CACTTCTTCATG-3ŌĆÖ; Bt1b, 5ŌĆÖ-GACGAGATCGTTCATGTT-GAACTC-3ŌĆÖ) (
Glass and Donaldson, 1995;
White et al., 1990). The PCR cycling conditions for the ITS region were 94┬░C for 4 min and 35 cycles of 94┬░C (35 sec), 52┬░C (55 sec), 72┬░C (1 min), and 72┬░C for 10 min. The PCR cycling conditions for the
╬▓tub gene were 98┬░C for 3 min and 30 cycles of 98┬░C (10 sec), 67┬░C (15 sec), 72┬░C (2 min), and 72┬░C for 10 min. The PCR reactions contained 0.5 ╬╝l of each primer, 0.5 ╬╝l of Taq DNA polymerase (Bioneer, Daejeon, Korea), 0.5 ╬╝l of each dNTP, 2.5 ╬╝l of 10├Ś PCR reaction buffer, 18.5 ╬╝l of distilled water, and 2.0 ╬╝l of template DNA (total volume 25 ╬╝l). The DNA concentrations were estimated on a 1% agarose gel by com-paring the PCR product band intensity with a 100 bp DNA ladder. DNA sequencing was performed by SolGent Com. Ltd. (Daejeon, Korea), and the nucleotide sequences were searched using the GenBank database BLASTn tool (
http://www.ncbi.nlm.nih.gov/BLAST/). Phylogenetic analysis was performed in MEGA7 by the neighbor-joining method (
Kumar et al., 2016;
Saitou and Nei, 1987).
BLAST analysis returned a 571 bp ITS-5.8S rDNA sequence and a 463 bp
╬▓tub sequence. The fungus identity was con-firmed; the analysis showed 100% similarity to the
S. sclerotiorum isolate CXL14041906 ITS-5.8S rDNA sequence (GenBank accession no. KX781301) and 100% similarity to the
S. sclerotiorum SS1 isolate KF545202
╬▓tub sequence. The representative isolate (GPSS003) sequences were deposited in the NCBI database (GenBank accession no. MG931017 for ITS-5.8S rDNA; accession no. MG931018 for
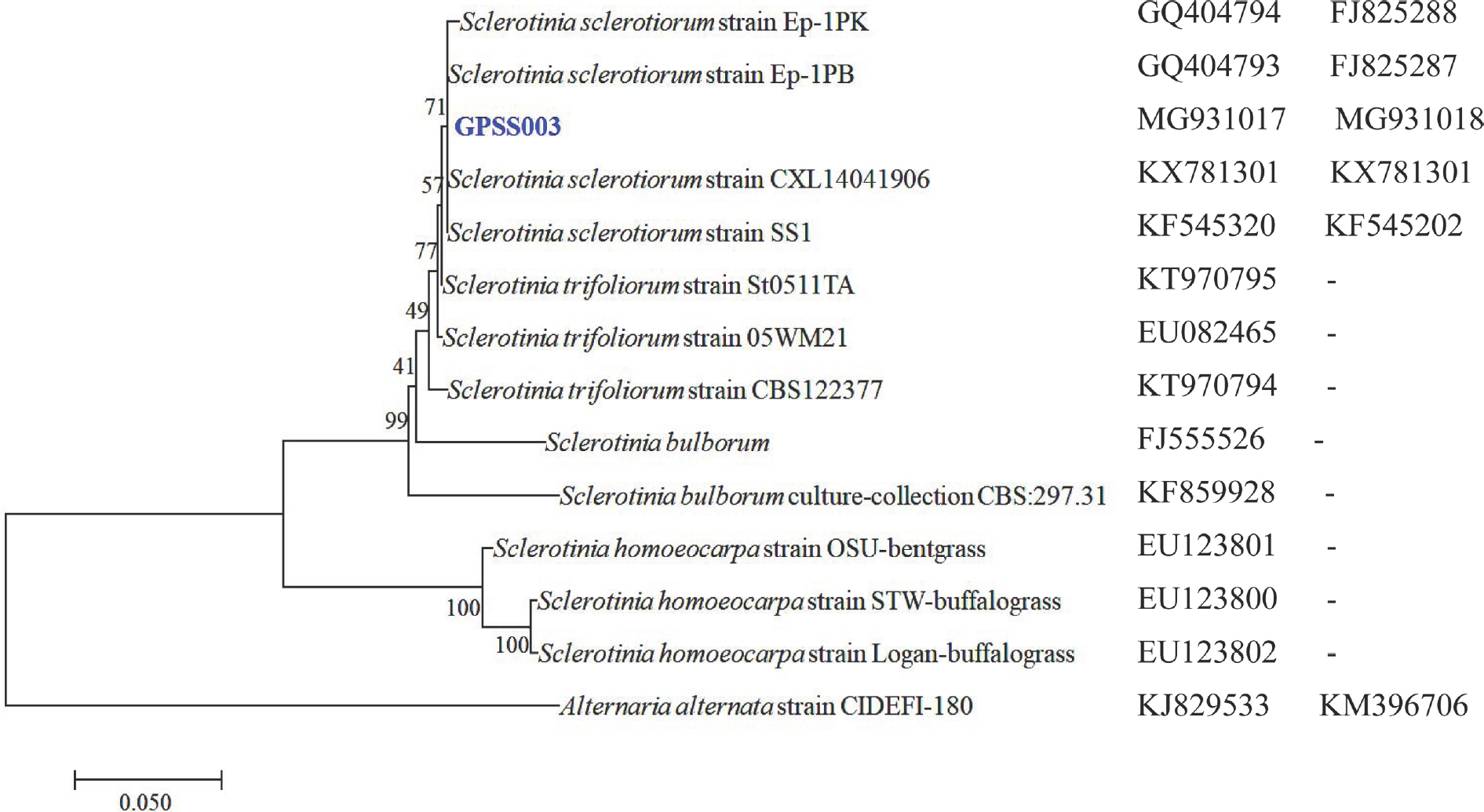
╬▓tub). The phylogenetic tree results (based on the combined ITS-5.8SrDNA and
╬▓tub sequences) placed the representative isolate within a clade that includes the
S. sclerotiorum reference isolates (
Fig. 2).
Fig.┬Ā2.
Phylogenetic analysis of Sclerotinia sclerotiorum isolate GPSS003 constructed using the neighbor-joining method based on the combined internal transcribed spacer and ╬▓tub gene sequence data. Alternaria alternata was the outgroup. The node numbers indicate the bootstrap values (1,000 replicates). The scale bar indicates the number of nucleotide substitutions.
To determine fungal pathogenicity, six green pea pods were surface-sterilized (1% NaOCl), rinsed with sterile distilled water (3├Ś), and dried (at RT). The sterilized pods were placed on moist filter paper in Petri dishes (20├Ś20├Ś5 cm). Three fungal mycelial discs (6.5 mm) were taken from the margins of actively growing 7-day-old colonies and placed into three sterilized pods. PDA discs were placed into three sterilized pods as a control. The Petri dishes were incubated in a growth chamber for 7 days at RT and 90┬▒10% relative humidity (
Han et al., 2013;
Kim et al., 2014). White mycelia (a symptom of rot) were seen on the inoculated pea pods after 5 days. The fungal pathogen was re-isolated from the diseased lesions on the inoculated pods and exhibited the same morphological characteristics as the original isolates (
Fig. 1B). Therefore, the fungal pathogen fulfilled Koch's pos-tulates, and
S. sclerotiorum was identified as the post-harvest green pea pod rot causal agent.
Morphological characterization, pathogenicity test, and phylogenetic analysis of the ITS region and
╬▓tub gene identified
S. sclerotiorum as the pathogen isolated from the green peas. In Korea, green pea field rot from
Sclerotinia infections has been documented, but the post-harvest disease has not (
Chang and Kim, 2003;
Kim et al., 2006).
S. sclerotiorum-induced stem and pod field rot have been reported in, e.g., Canada (
Farr and Rossman, 2021), Bangladesh (
Islam et al., 2020), and Korea (
Kim et al., 2006) but this is the first report of a post-harvest green pea pod rot in Korea. Post-harvest rotting caused by
S. sclerotiorum can lead to complete product loss, making it an economically important issue. These findings suggest that
Sclerotinia-induced rot may pose a threat to the long-term storage of produce, and an effective control strategy is needed.
Acknowledgments
This study was supported by a grant from the Rural Development Administration (project no. PJ012482), Republic of Korea. This study was also supported by the 2020 postdoctoral Fellowship Program (M. Aktaruzzaman) to the National Institute of Agricultural Sciences, RDA, Republic of Korea.
References
Boland, G. J. and Hall, R. 1994. Index of plant hosts of
Sclerotinia sclerotiorum.
Can. J. Plant Pathol. 16: 93-108.

Bolton, M. D., Thomma, B. P. H. J. and Nelson, B. D. 2005.
Sclerotinia sclerotiorum (Lib.) de Bary: biology and molecular traits of a cosmopolitan pathogen.
Mol. Plant Pathol. 7: 1-16.

Chang, S. W. and Kim, S. K. 2003. First report of Sclerotinia rot caused by
Sclerotinia sclerotiorum on some vegetable crops in Korea.
Plant Pathol. J. 19: 79-84.

Glass, N. L. and Donaldson, G. C. 1995. Development of primer sets designed for use with the PCR to amplify conserved genes from filamentous Ascomycetes.
Appl. Environ. Microbiol. 61: 1323-1330.



Han, K. S., Kim, J. Y., Park, J. H. and Shin, H. D. 2013. First report of
Sclerotinia stem rot of anemone caused by
Sclerotinia sclerotiorum in Korea.
Plant Dis. 97: 997.

Huang, H. C. and Kozub, G. C. 1989. A simple method for production of apothecia from sclerotia of Sclerotinia sclerotiorum
. Plant Prot. Bull. Taiwan 31: 333-345.
Islam, M. R., Prova, A., Akanda, A. M. and Hossain, M. M. 2020. First report of white mould caused by
Sclerotinia sclerotiorum on pea in Bangladesh.
J. Plant Pathol. 102: 941.

Kim, J.-Y., Aktaruzzaman, M., Afroz, T., Hahm, Y.-I. and Kim, B.-S. 2014. The first report of postharvest stem rot of kohlrabi caused by
Sclerotinia sclerotiorum in Korea.
Mycobiology 42: 409-411.



Kim, W. G., Hong, S. K. and Lee, S. Y. 2006. Occurrence of Sclerotinia rot in four leguminous crop caused by. Sclerotinia sclerotiorum. Plant Pathol. J. 22: 16-20.
Kim, W. G., Koo, H. M., Kim, K. H., Hyun, I. H., Hong, S. K., Cha, J. S. et al. 2009. List of Plant Diseases in Korea. 5th ed. Korean Society of Plant Pathology, Anyang, Korea. 853 pp. (In Korean)
Kohn, L. M. 1979. A monographic revision of the genus Sclerotinia. Mycotaxon 9: 365-444.
Kumar, S., Stecher, G. and Tamura, K. 2016. MEGA7: Molecular Evolutionary Genetics Analysis version 7.0 for bigger datasets.
Mol. Biol. Evol. 33: 1870-1874.



Naito, S. and Sugimoto, T. 1986. Sclerotinia stalk rot of sugar beets.
Ann. Phytopathol. Soc. Jpn. 52: 217-224.

Saitou, N. and Nei, M. 1987. The neighbor-joining method: a new method for reconstructing phylogenetic trees.
Mol. Biol. Evol. 4: 406-425.

Smith, E. A. and Boland, G. J. 1989. A reliable method for production and maintenance of germinated sclerotia of
Sclerotinia sclerotiorum.
Can. J. Plant Pathol. 11: 45-48.

White, T. J., Bruns, T., Lee, S. and Taylor, J. 1990. Amplification and direct sequencing of fungal ribosomal RNA genes for phyloge-netics. In: PCR Protocols: A Guide to Methods and Applications., eds. by M. A. Innis, D. H. Gelfand, J. J. Sninsky and T. J. White, pp. 315-322. Academic Press, San Diego, CA, USA.

Willetts, H. J. and Wong, J. A. L. 1980. The biology of
Sclerotinia sclerotiorum, S. trifoliorum, and
S. minor with emphasis on specific nomenclature.
Bot. Rev. 46: 101-165.






 PDF Links
PDF Links PubReader
PubReader ePub Link
ePub Link Full text via DOI
Full text via DOI Download Citation
Download Citation Print
Print






