Survey of the Incidence of Viral Infections in Calanthe spp. and Characterization of a GW Isolate of Cymbidium mosaic virus in Korea
Article information
Abstract
Cymbidium mosaic virus (CymMV) is a major virus infecting orchid plants and causing economic loss. In this study, the incidence of viral infection in Calanthe spp. at the Korean Institute of Calanthe was investigated using reverse transcription polymerase chain reaction. The CymMV infection rate was 42%, and the two viruses Odontoglossum ringspot virus and Cucumber mosaic virus had frequencies of 8% and 2%, respectively. Additionally, we characterized an isolate of CymMV, CymMV-GW, using biological tests and examined the nucleotide sequence properties of its complete genome. CymMV-GW induced chlorotic ringspots and chlorotic spot symptoms in inoculated leaves of Chenopodium amaranticolor and Nicotiana benthamiana, respectively. In this study, we have for the first complete genome sequence of CymMV-GW in Korea. The CymMV-GW genome was 6,225 nucleotides in length, excluding the poly-(A) tail, and showed whole-genome nucleotide and amino acid sequence identities of 97.7% and 100%, respectively, with the NJ-1 isolate of CymMV. Here, we report the complete genome sequence of the CymMV-GW isolate and viral infection rates for Calanthe spp. in Korea.
Introduction
Calanthe plants are distributed across cool and subtropical regions of Africa (Madagascar), Asia (Korea, Japan, China, Taiwan, India, and Nepal), the Indian Ocean, Australia (New Guinea), and the southwest Pacific, and approximately 29 species are known to occur in Asia (Gale and Drinkell, 2007; Govaerts et al., 2007). In Korea, Calanthe is narrowly distributed in Ulleungdo (Gyeongsangbuk-do), Anmyeondo (Chungcheongnam-do), Sinan, Jindo, Wando (Jeollanam-do), and the Jeju islands (Lee, 2011). The genus Calanthe includes approximately 171 to 200 species (Godo et al., 2010; Kurzweil, 2007; Suetsugu and Fukushima, 2014), but only five species (Calanthe discolor, C. bicolor, C. sieboldii, C. coreana, and C. reflexa) are known to occur in Korea (Lee and Kwack, 1983; So et al., 2012). Calanthe, similar to other orchid genera, is subject to infection by several plant viruses. So far, four viruses (Bean yellow mosaic virus, BYMV; Cucumber mosaic virus, CMV; Cymbidium mosaic virus, CymMV; and Odontoglossum ringspot virus, ORSV) have been reported in Calanthe spp. in Korea. Of these, CymMV (genus: Potexvirus, family: Flexiviridae) is the most prevalent and has caused significant economic losses (Wong et al., 1994; Zettler et al., 1990). CymMV has a monopartite, positive single-strand RNA genome that consists of five open reading frames (ORFs) containing genes encoding putative RNA-dependent RNA polymerase (RdRp), triple gene block (TGB) proteins, and coat protein (CP) (Wong et al., 1997). Some plant viruses containing the TGB movement protein are used to confirm cell-to-cell movement via protein interactions (Chou et al., 2013; Lim et al., 2008). CymMV has been used to study floral gene functions and lifespan using virus-induced gene silencing as a tool (Hsieh et al., 2013). For virus detection, various diagnostic methods have been developed, such as enzyme-linked immunosorbent assays (Vejaratpimol et al., 1998), polymerase chain reaction (PCR) (Lim et al., 1993), reverse transcription (RT)-PCR (Seoh et al., 1998), real-time PCR (Eun et al., 2000), and RT loop-mediated isothermal amplification assays (Lee et al., 2011).
Nevertheless, the virus has been studied with respect to its genetic diversity, homogeneity, relationships, distribution, phenotype, morphological characteristics, and taxonomy in Korea (Cho et al., 2007; Chung et al., 2014; Hyun et al., 1999; Kim et al., 2013; Lee et al., 2008; Srikanth et al., 2012). Although Jang (1998) studied viral disease in many kinds of orchids, including Calanthe, these studies were limited to an electron microscope analysis, biological tests, and diagnoses. Thus, we report the complete genome sequence and characterization of CymMV isolated from a Calanthe plant in Korea and the incidence of viral infections caused by different viruses at the Korean Institute of Calanthe.
Materials and Methods
Sample collection
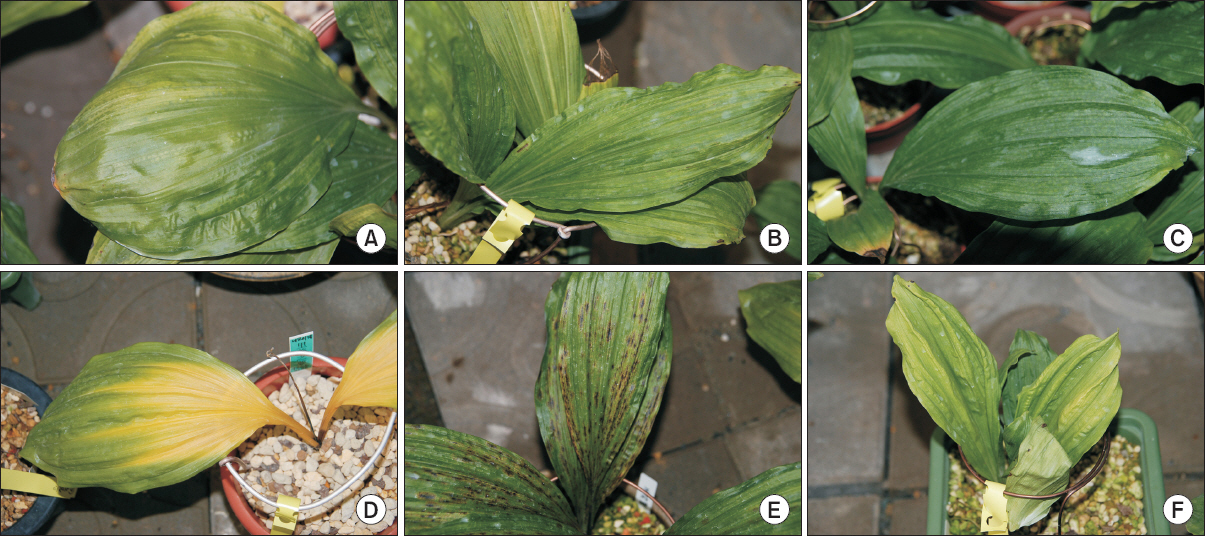
From 2012 to 2013, Calanthe spp. samples were provided by the Korean Institute of Calanthe (Gwangju, Korea). All 35 samples showed typical symptoms of viral infection, such as mild mosaic, mosaic, and malformation (Fig. 1), and were named GW isolates. The leaves with these symptoms were cut using a razor and the samples stored in a deep freezer (–80°C).
Virus diagnosis
In order to identify the infecting virus, total RNA was extracted from 35 samples of Calanthe spp. using Tri-Reagent (MRC Reagent, Cincinnati, OH, USA) following the manufacturer's instructions. The cDNA was synthesized using Oligo (dT6) primers with SuperScript™ II Reverse Transcriptase (Invitrogen, Carlsbad, CA, USA) according to the manufacturer's protocol. All samples were tested for eight viruses (BYMV, CMV, CymMV, ORSV, Clover yellow mosaic virus [ClYMV], Clover yellow vein virus [ClYVV], Turnip mosaic virus [TuMV], and Orchid fleck virus [OFV]) using specific primer sets by PCR with Elongase® Enzyme Mix (Invitrogen) (Table 1). The amplification conditions were as follows: an initial denaturation at 94°C at 2 minutes, 26 cycles of 94°C for 30 seconds, 55°C for 30 seconds, and 68°C for 1 minute, and a final extension at 68°C for 5 minutes. Amplified PCR products were confirmed by 1.0% agarose gel electrophoresis with ethidium bromide.
Biological test
To obtain a biological and molecular characterization of CymMV, a singly infected GW isolate was used (No. 1). The host range for CymMV-GW was determined by mechanical transmission with indicator plants for 34 species representing 7 families (Aizoaceae, Amaranthaceae, Brassicaceae, Chenopodiaceae, Cucurbitaceae, Leguminosae, and Solanaceae). Sap extracted from symptomatic leaf samples was used as the inoculum with 0.1 M potassium phosphate (pH 7.0), and tests were repeated three times per indicator plant. All indicator plants were maintained at 25°C–30°C in a greenhouse. To confirm local or systemic infections, at 8 weeks post-inoculation, inoculated and non-inoculated leaf samples were harvested and examined by RT-PCR as described above.
Complete genome sequencing and genome analysis
To determine the complete genome sequence, six primer sets were designed using the nucleotide sequences of CymMV isolates (GenBank accession Nos. HQ681906, EF125180, EF125179, AF016914, and JQ860108) from National Center for Biotechnology Information (NCBI) (Table 2). PCR products with expected sizes were obtained from CymMV-GW, and these products were cloned into a TA cloning vector (RBC Bioscience, Taipei, Taiwan) and sequenced (Solgent, Daejeon, Korea). To confirm the 5´- and 3´-terminal regions, extracted total RNA was subjected to rapid amplification of cDNA ends. First-strand cDNA was synthesized using primers (CymMV-R0, 5´- GAT TGA CTG TCA CGC CGA GA -3´) and (Oligo-dT15-Adaptor primer, 5´- GAC CAC GCG TAT CGA TGT CGA T16 -3´). The cDNA was purified using the ExgeneTM Plant SV Kit (GeneALL, Seoul, Korea). At the 5´ ends, a poly-(G) tail was added using purified cDNA with terminal deoxynucleotidyl transferase (Takara, Shiga, Japan). Each cDNA was amplified by PCR with primers (Oligo-dCD, 5´- GAC CAC GCG TAT CGA TGT CGA C17D -3´ and CymMV-R0) and (CymMV-F0, 5´- AGC CTG CTG AAT GGC AGC GC -3´ and Adaptor primer). PCR products were purified, cloned, and sequenced.

Primer sets used to amplify the complete genome of Cymbidium mosaic virus (CymMV) isolated from Calanthe spp.
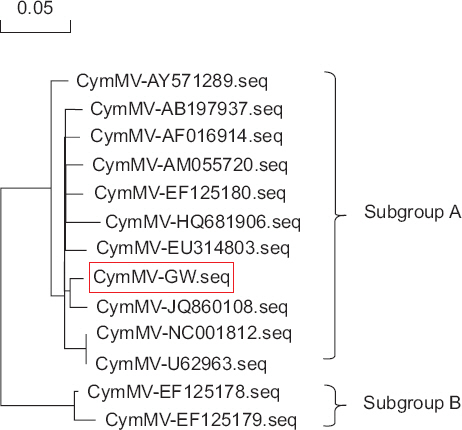
Phylogenetic analyses were conducted to examine the relationships among isolates with in the species CymMV, and to identify the most closely related isolates using DNAMAN software ver. 5.2.10 (Lynnon Biosoft, Quebec, QC, Canada). Sequence data for 12 CymMV isolates were obtained from NCBI. Multiple alignments were generated using complete genome sequences, and a phylogenetic tree was constructed using the maximum likelihood technique. Bootstrapping was performed using 1,000 replicates.
Results
To identify the infecting virus, all 35 collected samples were tested using RT-PCR. Among them, 15 were positive for CymMV, 3 were infected with ORSV, and 1 was infected with CMV. Two samples, GW isolate numbers 19 and 21, were co-infected with CymMV and ORSV. Five viruses (BYMV, ClYMV, ClYVV, TuMV, and OFV) were not detected in this test (Table 3).
To confirm the host range of CymMV GW isolate, the bioassay carried out using various test plants. After three weeks, chlorotic ringspots were detected on inoculated leaves of Chenopodium amaranticolor. Nicotiana benthamiana showed chlorotic spots on inoculated leaves at two weeks post-inoculation. N. glutinosa produced mosaic symptoms on non-inoculated leaves. N. tabacum var. Turkish and N. tabacum cv. Samsun showed malformation symptoms on upper leaves (Fig. 2). The remainder of the test plants is without symptoms. Two species (C. amaranticolor and N. benthamiana) yielded RT-PCR products consistent with the expected size of CymMV, while the virus was not detected for 32 species whose indicator plants showed symptoms of viral infection.
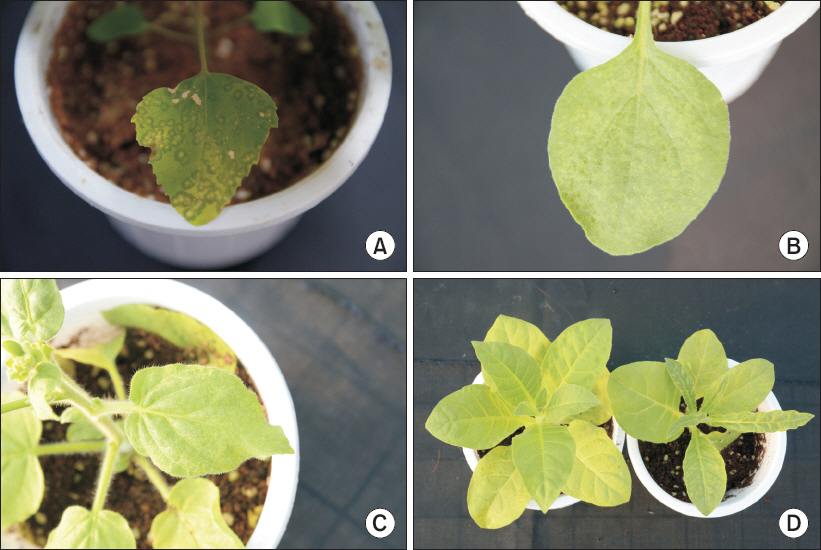

Mechanical inoculation of Cymbidium mosaic virus from Calanthe plants to several indicator plants. (A) Chenopodium amaranticolor produced chlorotic ringspot symptoms on inoculated leaves. (B) Nicotiana benthamiana showed chlorotic spots on inoculated leaves. (C) Systemic mosaic symptoms in N. glutinosa. (D) Left: healthy plant, right: malformation symptoms on upper leaves in N. tabacum var. Turkish.
The genome of the CymMV-GW isolate consisted of 6,225 nucleotides (nt), excluding the poly-(A) tail. The virus contained five ORFs, ORF1 (RdRp, nucleotide positions 73–4,326), ORF2 (TGB1, 4,332–5,033), ORF3 (TGB2, 5,008–5,346), ORF4 (TGB3, 5,201–5,476), and ORF5 (CP, 5,479–6,150). The 5´ and 3´ untranslated regions were 72 and 75 nt long, respectively (Fig. 3). CymMV-GW was most closely related to the CymMV NJ-1 isolate, which was isolated from Phalaenopsis spp. in China (GenBank accession No. JQ860108). According to a previous study by Vaughan et al. (2008), the GW isolate clustered into a group referred to as subgroup A (Fig. 4). The complete nucleotide sequence of CymMV-GW showed the highest identity with the NJ-1 isolate (nt identity, 97.7%; amino acid identity, 100%). Other CymMV isolates shared more than 95% identity at the nucleotide and amino acid levels, except for two USA isolates. The CymMV 18-1 and 18-30 isolates (GenBank accession Nos. EF125178 and EF125179), which were isolated from Dendrobium spp. in the United States, showed nucleotide sequence identities of 88.7% and 87.5%, respectively, with CymMV-GW (data not shown).
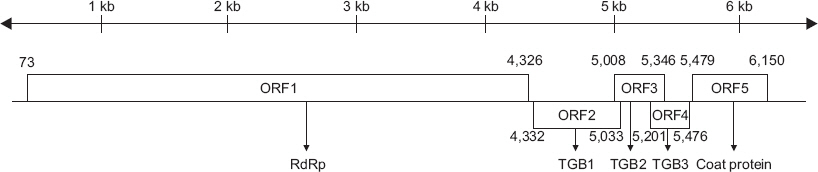
Schematic diagram of the complete genome organization of Cymbidium mosaic virus (CymMV). CymMV contains five open reading frames (ORFs): ORF1 (RNA-dependent RNA polymerase, nucleotide positions 73–4,326), ORF2 (triple gene block 1 [TGB1], 4,332–5,033), ORF3 (TGB2, 5,008–5,346), ORF4 (TGB3, 5,201–5,476), and ORF5 (coat protein, 5,479–6,150). RdRp, RNA-dependent RNA polymerase.
Discussion
Calanthe spp. are expected to be new economically important horticultural crops. A great deal of research has been carried out to develop export-quality crops, but viral disease is a major cause of low quality. Here, we surveyed cultivated Calanthe plants in Korea and obtained 35 samples to test for the presence of the viruses BYMV, ClYVV, ClYMV, CMV, CymMV, OFV, ORSV, and TuMV. CymMV had the highest infection rate (42%) among the eight viruses (BYMV: 0%, ClYVV: 0%, ClYMV: 0%, CMV: 2%, OFV: 0%, ORSV: 8%, and TuMV: 0%). These results imply that CymMV is a major virus infecting Calanthe plants, similar to other orchid plants (Hu et al., 1993).
In the host range test, among the 34 species examined, symptoms of viral infection were observed in 6 species (C. amaranticolor, N. benthamiana, N. glutinosa, N. rustica, N. tabacum cv. Samsun, and N. tabacum var. Turkish). RT-PCR results for C. amaranticolor and N. benthamiana were positive for CymMV, but four indicator plants tested negative for the virus. Additionally, harvested indicator samples were tested using other virus-specific primers (Table 1) and the results were negative. The present results suggest that identification of viruses on undetected samples is required. According to Moles et al. (2007), the CP and RdRp genes can be used to distinguish a subgroup from other CymMV isolates. Thus, we constructed a phylogenetic tree using 50 reported CP gene sequences of CymMV obtained from the NCBI database. Based on the phylogeny, the CP sequences could be divided into two subgroups, and GW isolates clustered in subgroup A (data not shown).
In this study, the complete genome sequence of CymMV isolated from Calanthe species in Korea was determined for the first time and the infection rate and host range of CymMV were confirmed. These data will facilitate the development of virus-induced gene silencing tools, expression vectors, and breeding systems.
Acknowledgement
This research was supported by a grant from the Animal and Plant Quarantine Agency (QIA), Ministry of Agriculture, Republic of Korea (Project Code No. Z-1542051-2013-15-01).